Research
COMPREHENSIVE PROTEOMICS
Proteins are a cell’s machinery and perform most functions within living organisms. Many illnesses and diseases are caused by problems that occur at the protein level and understanding what goes wrong at the protein level will help future disease prevention, diagnosis, and treatment. Proteomics seeks to understand the role of proteins in living systems and begins with the systematic identification and quantitation of as many proteins as possible from a cell, tissue, organism, or biological community (e.g. a microbiome) or other sample. A goal of comprehensive proteomics is to measure all the proteins present in a sample, but this is currently not possible due to the complexity of the samples and limitations of measurement technologies. The factors that contribute to the complexity of proteomes are numerous while key factors that contribute to complexity include: the large number of proteins present in an organism at any point in time, protein concentrations changes with time, most proteins have numerous modification states, and the dynamic range of protein relative abundance is huge. We focus on the development of new microfluidic and nanofluidic systems to improve comprehensive proteome analysis for mass spectrometry (MS) based proteomics. A goal of our systems is to integrate sample preparation and separation into microfluidic/nanofluidic systems before introducing into the MS by electrospray ionization (ESI). Once fully developed, these systems will increase the number of proteins identified providing a more complete description of the proteome while increasing the speed and reproducibility.
NANOFLUIDIC/MICROFLUIDIC INTERFACES (NMIs)
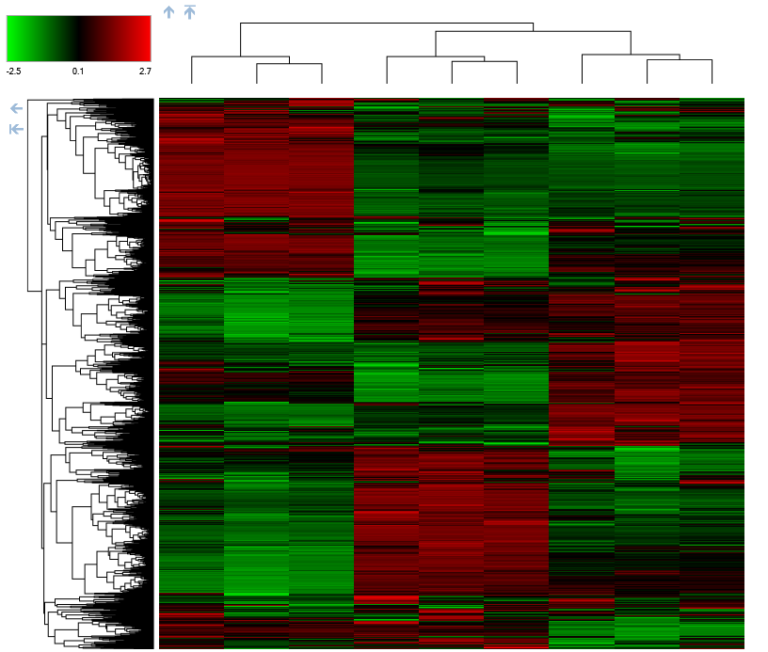
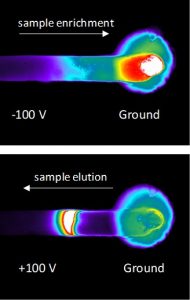
One functional module of these larger microfluidic systems are nanofluidic/microfluidic interfaces (NMIs) that provide rapid enrichment of proteins and peptides on the microfluidic chip. We have investigated the underlying fundamentals of transport within the NMIs to use them within our larger microfluidic system and ultimately improve the detection of proteins and their various proteoforms. An unique aspect of our NMI concentrators is that they enrich samples based on electrophoresis and are compatible with microchannel coatings that reduce surface charge and deleterious protein adsorption. In the images on the left, the enrichment of a sample that is not detectable at its initial concentration is shown in a microchannel terminated with a nanocapillary membrane. The negatively charged green fluorescent protein sample is enriched by electrophoresis, and an enriched band is eluted by switching the direction of the electric field. The images were acquired on an inverted fluorescent microscope, viewing the microfluidic device from the bottom, with white showing the highest protein concentration and blacking showing regions with near zero protein concentration.
LOW FREQUENCY AC ELECTROPHORESIS
Currently, medical diagnostic devices are being revolutionized as numerous biomolecular tests are being developed. These next-generation diagnostic systems will provide earlier detection, cost less, and operate in remote areas. Our lab is developing novel point-of-care diagnostic devices based on:
1) traveling wave electrophoresis,
2) non-linear electrophoresis of single particles,
3) mechanophenotyping of individual cells.
These systems will be used to detect pathogenic bacteria, and viruses in body fluids and water; and differentiate cells that are healthy, infected by bacteria, or infected by a virus.
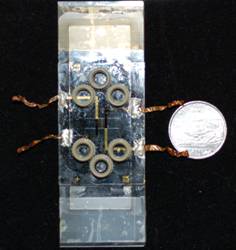


MICROFLUIDIC CHIP for CELL CULTURE
The PDMS made microfluidic chip can benifit the process of cell culture with easier experiment processing, co-culture up to 4 different cell lines, and provideing multi concentraiton treatment on one chip. Combing to the pumping system for culture medimum and drug treatment, four levels of concentraiton are generated consistently after the mixing process. The purposes of culturing cells in the microfluidic chip are to eliminate the experimental variables, simplify the experimental process, and simulate in vivo nutrient perfusion. In our lab, the application of on-chip cell culture is mainly focusing on cancer cell lines for multi hypothesis testing.
